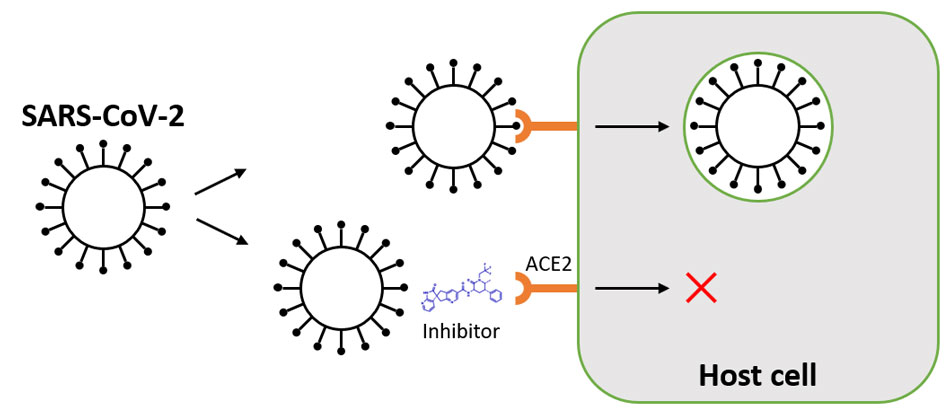
Eleven individuals from the Danish outbreak had a variant termed cluster 5, which had 3 additional mutations in spike (del69_70, I692V, and M1229I). Many SARS-CoV-2 sequences from the Netherlands and Danish outbreaks had a Y453F mutation in the receptor binding domain of spike, which might mediate increased binding affinity for mink ACE2 (angiotensin-converting enzyme 2). 7 In early November 2020 Danish authorities reported 214 cases of human coronavirus disease 2019 (COVID-19) associated with mink farms. 6 Genomic and epidemiologic investigation of an early outbreak in the Netherlands demonstrated human to mink, mink to mink, and mink to human transmission. 4, 5Ĭoncerning outbreaks of SARS-CoV-2 began to emerge on mink farms in the Netherlands and Denmark in late spring and early summer 2020. More recently, complementary work in animal models indicates that 614G viruses transmit more efficiently. 3 The effect size was modest, and the varying models did not always achieve statistical significance. A recent population genetic and phylodynamic analysis of more than 25 000 sequences from the UK found that viruses bearing 614G did appear to spread faster and seed larger phylogenetic clusters than viruses with 614D. This plausible null hypothesis led many in the evolution community to doubt that the D614G mutation was adaptive, despite in vitro data showing its effects on receptor binding. This raised the possibility that global dispersal of this mutation could have resulted from chance founder events, in which viruses harboring 614G just happened to initiate the majority of early transmission events in multiple locations. However, subsequent sequencing efforts identified the D614G mutation in viruses in several Chinese provinces in late January. This apparent convergent evolution was suggestive of natural selection and an adaptive benefit of D614G. 2 The mutation initially appeared to arise independently and simultaneously sweep across multiple geographic regions. The D614G mutation in the spike glycoprotein of SARS-CoV-2 was first detected at a significant level in early March 2020 and spread to global dominance over the next month. 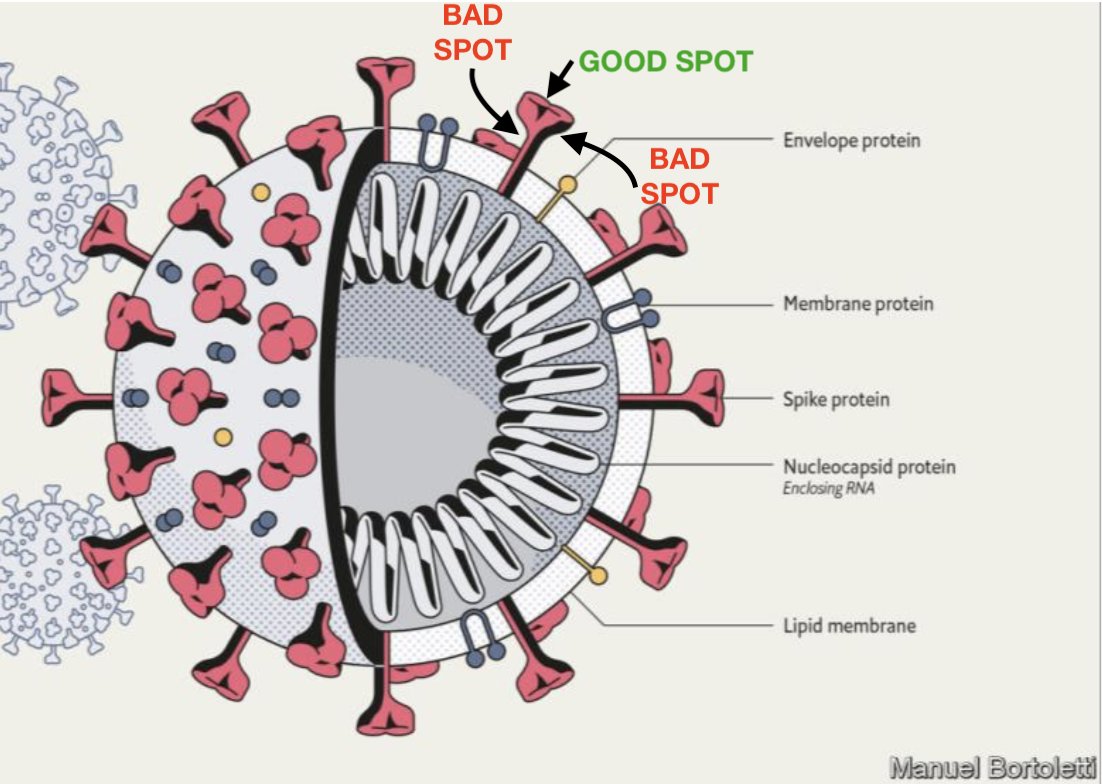
Strictly speaking, a variant is a strain when it has a demonstrably different phenotype (eg, a difference in antigenicity, transmissibility, or virulence).Įvaluation of a new SARS-CoV-2 variant should include assessment of the following questions: Did the variant achieve prominence through natural selection or chance events? If the evidence suggests natural selection, which mutation(s) are being selected? What is the adaptive benefit of these mutations? What effect do these mutations have on transmissibility and spread, antigenicity, or virulence? 
This term is somewhat less precise because 2 variants can differ by 1 mutation or many. Genomes that differ in sequence are often called variants. Mutation refers to the actual change in sequence: D614G is an aspartic acid-to-glycine substitution at position 614 of the spike glycoprotein. This same interplay of natural selection and chance events shapes virus evolution within hosts, in communities, and across countries.Īlthough the terms mutation, variant, and strain are often used interchangeably in describing the epidemiology of SARS-CoV-2, the distinctions are important. The mutations present in the genomes of these viral ancestors will dominate the population regardless of their effects on viral fitness. For example, a “founder effect” occurs when a limited number of individual viruses establish a new population during transmission. However, mutations can also increase and decrease in frequency due to chance events. Those that confer a competitive advantage with respect to viral replication, transmission, or escape from immunity will increase in frequency, and those that reduce viral fitness tend to be culled from the population of circulating viruses. In most cases, the fate of a newly arising mutation is determined by natural selection. Coronaviruses, however, make fewer mutations than most RNA viruses because they encode an enzyme that corrects some of the errors made during replication. 
1 RNA viruses typically have higher mutation rates than DNA viruses. Mutations arise as a natural by-product of viral replication. Shared Decision Making and Communication.Scientific Discovery and the Future of Medicine.Health Care Economics, Insurance, Payment.Clinical Implications of Basic Neuroscience.Challenges in Clinical Electrocardiography.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
